focimt is the stand-alone command line application for performing the seismic moment tensor inversion in time domain optimized for local-to-regional applications and seismic networks (Cartesian coordinate system is used). The core of the program is based on the formal description presented in Fitch et al. (1980) and Wiejacz (1992). The application uses first P-wave motion amplitudes and duration of the first P-wave ground velocity pulse to invert for the unconstrained full moment tensor. In addition, the constrained trace-null and double-couple moment tensors may be calculated as well. The calculated full moment tensor can be decomposed into isotropic (ISO) and deviatoric parts (CLVD/DC). The output information can be exported to a highly customized output text files. focimt is also capable to provide high-quality graphical representation of the moment tensor inversion in a form of beach-balls. The graphical representation can be exported to both raster and vector file formats such as PNG, PDF, PS and SVG. The precompiled binaries are available for both Windows/Linux platforms.
focimt command line tool is a part of hybridMT moment tensor inversion and refinement software package designed for MATLAB environment. The package contains wrapper function focimt.m that allows to conveniently handle the input parameters and inversion results directly in MATLAB. Numerous parameters can be specified to manage the input data, define seismic moment tensor inversion/optimization parameters as well as to handle the properties of graphical and text output. The package contains extensive MATLAB/PDF documentation with examples and tutorials. For details see the dedicated hybridMT webpage.
Authors
Grzegorz Kwiatek and Patricia Martínez-Garzón.
Referencing
When you find focimt application useful, please refer to it in your work:
Kwiatek, G., Martínez-Garzón, P. and M. Bohnhoff (2016). HybridMT: A MATLAB/Shell Environment Package for Seismic Moment Tensor Inversion and Refinement. Seismological Research Letters 87 (4), DOI: 10.1785/0220150251.
Download
focimt is part of hybridMT software package which contains an extensive documentation of focimt tool as well as the MATLAB wrapper for focimt, focimt.m. Please download the hybridMT package from the dedicated site to get the newest version of focimt tool.
Documentation
To get the list of all command line options of focimt command line tool, execute:
focimt -h
The detailed documentation of focimt command line options as well as examples of application are presented in [wpfilebase tag=fileurl id=35 linktext=’hybridMT software package manual’ /]. Below you will find couple of examples of application of the command line tool.
License
focimt is licensed under GNU General Public License (GPL)
Quick example
Using focimt command line or (equivalently focimt.m MATLAB routine bundled in hybridMT package) is very easy. Having the ASCII input file containing event and phase data, the moment tensor inversion may be started directly from the console window (or from MATLAB command prompt by starting the line with exclamation mark):
focimt -i example1.txt -s FTD -t PNG -p SU -z 300 -d M
The moment tensor inversion results in generation of output ASCII containing moment tensor inversion results and graphical representation of the solution in a form of
beach ball plot. The equivalent call from MATLAB command prompt using focimt.m routine would like as follows:
Solution = focimt('example1.txt','BeachBallFormat','PNG','Projection','schmidt','Hemisphere','upper','BeachBallSize',300);
resulting in output parameters being stored in output cell array Solution than can be easily further used in MATLAB environment.
Both calls generate graphical representation(s) of moment tensor solution as in the following picture:
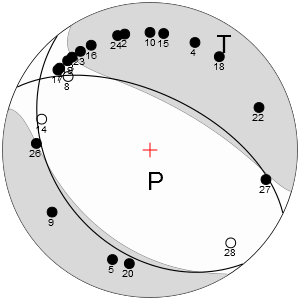
More examples on how to use focimt command line tool is presented the program documentation, which is a part of hybridMT package.
FAQ
-
What is the magnitude limit for application of fociMT or hybridMT algorithms?
The fociMT algorithm is point-source moment tensor inversion and the fault rupture processes (e.g. directivity or spatial changes in elementary moment tensors) are NOT considered. As such, the algorithm is valid for events with simple source time functions that can be represented by a single pulse. Therefore, it is suggested to use the algorithm for events with moment magnitudes not exceeding Mw 4.
-
Is fociMT algorithm based on full waveform inversion?
No. The input for moment tensor inversion using fociMT algorithm is area below first P-wave ground displacement pulse with sign information. As such, the inversion algorithm fall into the class of “amplitude MT inversion” algorithms (i.e. it is different from e.g. ISOLA algorithm)
-
What is the “omega” parameter in the focimt input file and how to calculate it?
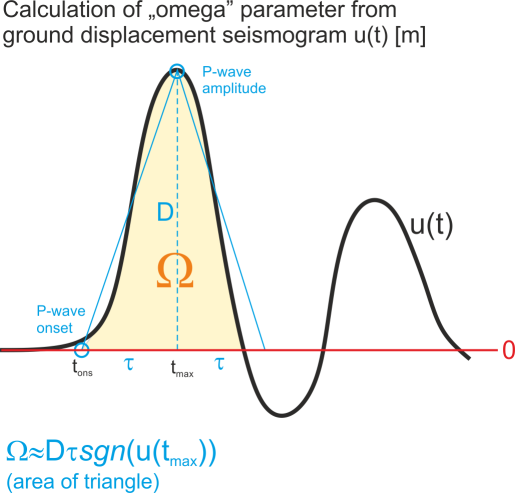 “omega” parameter which is a part of ASCII input file is basically the area below the first P-wave ground displacement pulse (in time domain) or equivalently the spectral level taken from the amplitude spectrum of ground displacement pulse. The unit of “omega” parameter is [m * s] (meter times second), and it comes from the fact that we multiply the amplitude of first P-wave pulse using ground displacement seismogram in [m] by the half of duration of the first P-wave pulse in [s]. Regardless of whether time or frequency domain was used to extract the “omega” value, the parameter should contain the sign information. The sign of “omega” is in accordance with the seismological convention, i.e. it is positive for the positive first P-wave ground displacement amplitude (i.e. ground motion AWAY from the seismic source) and negative otherwise (ground motion TOWARDS the seismic source).
“omega” parameter which is a part of ASCII input file is basically the area below the first P-wave ground displacement pulse (in time domain) or equivalently the spectral level taken from the amplitude spectrum of ground displacement pulse. The unit of “omega” parameter is [m * s] (meter times second), and it comes from the fact that we multiply the amplitude of first P-wave pulse using ground displacement seismogram in [m] by the half of duration of the first P-wave pulse in [s]. Regardless of whether time or frequency domain was used to extract the “omega” value, the parameter should contain the sign information. The sign of “omega” is in accordance with the seismological convention, i.e. it is positive for the positive first P-wave ground displacement amplitude (i.e. ground motion AWAY from the seismic source) and negative otherwise (ground motion TOWARDS the seismic source).In case of simple impulsive source time functions it is typically not necessary to calculate the integral of ground displacement first P-wave pulse. Instead, the amplitude D (cf. picture) including sign information of first P-wave pulse may be taken from ground displacement seismogram (the seismogram should be calibrated to [m]) and multiplied by the the rise time (i.e. time interval between the P-wave onset and P-wave amplitude, t_max-t_ons in seconds [s]). In this case, the integral of the pulse is simply replaced by the area of the triangle.
-
What full moment tensor decomposition scheme is used in focimt.m?
The MATLAB wrapper focimt.m has no option for changing the decomposition scheme, which is currently fixed to Jost and Herrmann’s (1989) approach. However, the focimt application has implemented an alternative decomposition scheme based on Vavrycuk’s (2001) approach. In focimt application this is specified by option -d D or -d Y (see the documentation of focimt application for details). Therefore, the hardcoded decomposition scheme in focimt.m may be changed from Jost and Herrmann’s to Vavrycuk’s approach simply by changing the following line from (note change from “D” to “Y” in option -d):
commandline = ['-i ' temp '.txt -d DWAFTUMVE -o ' temp ' -s ' solutions ' -n ' normfunc ' ' bbsize ' ' bbformat ' ' bbproj ' ' jacknife ' ' bootstrap ' ' vmodel ' ' ball];
to
commandline = ['-i ' temp '.txt -d YWAFTUMVE -o ' temp ' -s ' solutions ' -n ' normfunc ' ' bbsize ' ' bbformat ' ' bbproj ' ' jacknife ' ' bootstrap ' ' vmodel ' ' ball];
Future version of focimt.m implement the decomposition switch parameter.
- How should I provide takeoff angle in RAW ASCII file format?
-
What is the azimuth convention for RAW ASCII file format?
The azimuth of a vector from the seismic hypocenter to the seismic station is measured in degrees, and it is positive from north towards east. 0,90,180, and 270 degrees corresponds to the station located North, East, South, and West from the earthquake hypocenter.
-
What is the incidence angle convention for RAW ASCII input file?
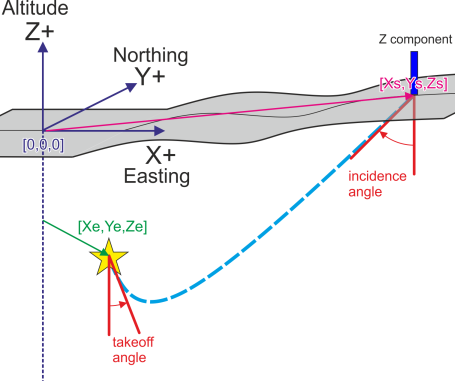 The angle of incidence in RAW ASCII input file follows the standard seismological convention. The angle of incidence to the sensor is measured in degrees from down direction (towards center of Earth). In other words, the seismic ray pointing directly up (perpendicular towards the surface) has 0 degrees incidence angle and pure horizontal ray (parallel to the surface) has 90 degrees incidence angle.
The angle of incidence in RAW ASCII input file follows the standard seismological convention. The angle of incidence to the sensor is measured in degrees from down direction (towards center of Earth). In other words, the seismic ray pointing directly up (perpendicular towards the surface) has 0 degrees incidence angle and pure horizontal ray (parallel to the surface) has 90 degrees incidence angle.Note #1: In case of pure horizontal ray (incidence = 90) one should not expect P-wave arrival to be visible on vertical component. Phases with incidence angle = 90 degrees should be disregarded from processing.
Note #2: Large values of incidence angle (>80 degrees) leads to high corrections for “omega” parameter, as the “omega” is divided by the cosine of the angle of incidence. This typically leads to overestimation of total and scalar seismic moment, as well as it may influence the moment tensor decomposition.
-
Why I get error message on “incorrect expression or statement” when running drawhudsonnet.m or drawstereonet.m?
If you get error similar to this:
>> drawhudsonnet; ??? Error: File: drawhudsonnet.m Line: 119 Column: 3 Expression or statement is incorrect--possibly unbalanced (, {, or [.it is likely due to the fact that your version of MATLAB does not support empty output arguments that are specified in MATLAB as “~” (tilde character). If possible, please update your MATLAB to the newest version that handles empty output arguments. Alternatively, please change every null output variable “~” to any valid and not used variable name, e.g. “dummy”:
[~,DD] = eig(M);
to
[dummy,DD] = eig(M);
This problem will be resolved in future version of hybridMT.
-
Why I receive error messages “Argument ‘###’ failed validation with error: ####”
Consider a scenario of executing the drawstereonet.m function:
[X,Y] = drawstereonet([10 20 30 40],[50 60 70 80]); ??? Error using ==> drawstereonet at 43 Argument 'AZM' failed validation with error: Undefined function or method 'ismatrix' for input arguments of type 'double'.
This error results from the fact your MATLAB version does not contain ismatrix() routine. If it is possible, please update the MATLAB to the newest version. Alternatively, you can try to disable the validation routine. In case of example above, by changing line:
p.addOptional('AZM', [], @(x) ismatrix(x) && isnumeric(x) );to:
p.addOptional('AZM', [], @(x) isnumeric(x) );
Acknowledgements
focimt uses portions code by Pawel Wiejacz (Institute of Geophysics, Polish Academy of Sciences, Warsaw, Poland) . focimt binaries uses CAIRO library (http://cairographics.org/). The source code also contains routines from PSMECA program, which is a part of Generic Mapping Tools (GMT) software package available to download from http://gmt.soest.hawaii.edu/. The 1D velocity model ray-tracing routines were translated from hypoDD v1.3 package by Felix Waldhauser, see http://www.ldeo.columbia.edu/~felixw/hypoDD.html for details.
The Authors wish to thank Pawel Wiejacz, Boguslaw Domański, Marek Burdzy, Grzegorz Makowski, Marian Król, Wojciech Dębski, Ilona Kwiatkowska, Marcin Pussak, Anita Zych-Kotwicka, Artur Kotwicki, Łukasz Rudziński, Grażyna Maziarz-Wiorek, Nikos Melis, Beata Orlecka-Sikora, Bożena Hersztowska, Katrin Plenkers, Oliver Germer, Grzegorz Lizurek, Wojciech Białoń, Maria Kozłowska, Konstantinos Leptokaropoulos, Joanna Kocot, Jerzy Giza, Krystyna Stec for help, feedback and beta-testing of the software.
References
Fitch, T. J., D. W. McCowan, and M. W. Shields (1980). Estimation of seismic moment tensor from teleseismic body wave data with application to intraplate and mantle earthquakes, J. Geophys. Res. 85, 3817–3828.
Knopoff, L., and M. J. Randall (1970). The compensated linear-vector dipole. A possible mechanism for deep earthquakes, J. Geophys. Res. 75, 1957–1963.
Wiejacz, P. (1992). Badanie mechanizmów wstrząsów górniczych przy wykorzystaniu tensora momentu sejsmicznego, Institute of Geophysics, Polish Academy of Sciences.